By Tammie Tam, Molecular and Medical Microbiology ‘22
Humanity has always been intrigued with the possibility of life existing elsewhere in the universe. In 1977, NASA attached the Golden Record, a detailed account of humanity and Earth, onto the Voyager 1 and 2 space probes [1]. They intended for the records to be a way to share human knowledge with intelligent life forms that may stumble upon the space probes in interstellar space. While the Voyager 1 and 2 entered interstellar space in 2012 and 2018, they have yet to encounter any intelligent life forms capable of deciphering the record [2]. But extraterrestrial life doesn’t have to be “intelligent.” Life can be as simple as a swimming bacteria.
With the broad possibility of what life can look like, most scientists rely on what they understand about life’s limits on Earth to narrow down possible markers of extraterrestrial life. In 2022, a team led by Sara Walker at Arizona State University defined the types of biochemical functions likely to be found sustaining life throughout the universe [3]. Their work is a step forward in predicting how life may exist in places that look nothing like Earth by identifying potential signs of life using chemistry.
Walker’s team did this by computing scaling laws, which represent the proportional relationship between two quantifiable variables by some power factor [3]. Often these laws are used to demonstrate the universality of phenomenons found in physics. A simple example is the relationship between the surface area and volume of a cube. Given two cubes of differing sizes, the ratio of the surface areas is equivalent to the ratio of the volumes by a power or scaling exponent of ½ [4]. What makes this a universal principle is that no matter what kind of material the cube is made of, or where the cube is found, the relationship between surface area and volume stays the same.
In biology, scaling laws are not as commonly applied, but certain biological characteristics have still been shown to be able to scale to each other. For example, scientists have defined scaling laws between body mass and other features like growth rates, metabolic rates, and life spans [5]. These scaling laws in biology are typically defined by power laws: y = axk, where y is the rate of one variable, a is a constant, x is the rate of the other variable, and k is the scaling exponent [5]. Scaling laws in no shape or form provide the mechanism of how one feature regulates another feature and vice versa. Instead, they show that no matter the differences in detail between systems of different organisms, there is an underlying commonality between them all.
One caveat is that scaling laws for these well-studied biological features like growth and mass may only apply to life forms on Earth. For instance, while the final body mass and maximum growth rate within and across many taxonomic groups generally scale with each other at k = ¾, there’s no reason that this growth scale must remain the same across extraterrestrial life [5]. Because there are many factors that contribute to growth that scaling laws don’t reveal, the growth scale may be influenced by some adaptation to living on Earth. This is evident by the fact that certain taxonomic groups don’t follow the same scaling patterns in terms of growth rate and mass. Thus, scaling laws in biology are limited in their application when examining certain biological features.
However, chemistry throughout the universe is bound by the same laws and would not change whether on Earth or Mars or an asteroid in another galaxy. Walker’s group, therefore, started thinking about how, at its core, life is run by different chemical reactions [3]. Since atoms and chemicals are universal, perhaps the chemistry of life on Earth, or biochemistry, can be generalized to include extraterrestrial life.
The group studied enzymes, which are proteins that help chemical reactions run in the body. They collated databases on all known enzymes in the tree of life. The enzyme’s associated classes, reactions, and components are systematically encoded by a unique Enzyme Commission numerical identifier. They used the ID numbers to group enzymes into classes, such as oxidoreductases, transferases, hydrolases, lyases, isomerases, and ligases. ID numbers also helped them count the unique enzymes in each class [3].
By computing scaling laws as a statistical measure, they found that the number of unique enzymes within any particular class increases as the total number of enzymes from all classes increases [3]. This scaling relationship can generally be defined by y = axk, with k > 0 for all enzyme classes.
While each enzyme class demonstrated different scaling behaviors, what matters for determining the universality of a biochemical function of an enzyme class to have the same scaling behavior between all domains of life: archaea, bacteria, and eukaryotes. In their study, they found each enzyme class except for lyases to have scaling exponents within 95% confidence interval between domains, meaning the scaling behavior is similar enough to establish all enzyme classes except for lyases as universal biochemical functions [3].
The issue with lyases is resolved when lyases are grouped with hydrolases. Both enzyme classes deal with breaking down molecules. Together, their scaling behaviors between domains of life fall within a 95% confidence interval of each other, meaning lyases may potentially also constitute a universal biochemical function if reclassified with hydrolases [3].
These results reveal how these enzyme classes can inform what biochemical functions allow life to occur regardless of specific chemical components. The group reasoned that since they did not examine at the level of mechanistic details of how enzymes function, this work can be more generalizable beyond the tree of life as we know it [3].
This research will motivate others across various disciplines to propose new ideas to predict features of life outside of the immediate solar system. And until we find evidence of life (or until they find us), there are still many more ways scientists can try to bring some understanding to the unknown.
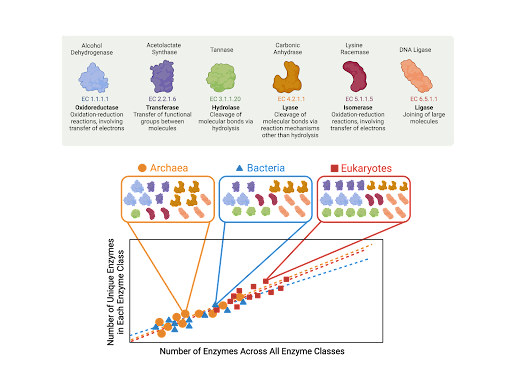
Figure 1. Here shows a conceptual schematic of how the number of unique enzymes in each enzyme class relate to the total number of enzymes across all enzyme classes, and how these scaling relationships compare between archaea, bacteria, and eukaryotes. The enzyme classes examined are oxidoreductase (EC 1), transferase (EC 2), hydrolase (EC 3), lyase (EC 4), isomerase (EC 5), or ligase (EC 6).
References:
- Voyager – what’s on the golden record. [accessed 2022c Apr 26]. https://voyager.jpl.nasa.gov/golden-record/whats-on-the-record/.
- Voyager – mission status. [accessed 2022b Apr 26]. https://voyager.jpl.nasa.gov/mission/status/.
- Gagler DC, Karas B, Kempes CP, Malloy J, Mierzejewski V, Goldman AD, Kim H, Walker SI. 2022. Scaling laws in enzyme function reveal a new kind of biochemical universality. Proc Natl Acad Sci USA. 119(9):e2106655119. doi:10.1073/pnas.2106655119. [accessed 2022 Apr 26]. https://pnas.org/doi/full/10.1073/pnas.2106655119.
- Introduction to scaling laws. [accessed 2022a Apr 26]. https://www.av8n.com/physics/scaling.htm.
- Hatton IA, Dobson AP, Storch D, Galbraith ED, Loreau M. 2019. Linking scaling laws across eukaryotes. Proc Natl Acad Sci USA. 116(43):21616–21622. doi:10.1073/pnas.1900492116. [accessed 2022 Apr 26]. https://pnas.org/doi/full/10.1073/pnas.1900492116.

