By Lia Freed-Doerr, Cognitive Science, Neurobiology, Physiology & Behavior ‘22
Author’s Note: In fall quarter, I got into contact with the Tian lab in the Department of Biochemistry and Molecular Medicine in order to learn more about optogenetic techniques and the difficulties of in vivo sensing of neural dynamics and, with the mentorship of a postdoctoral researcher, I have learned more about different high-resolution sensors (or indicators) and expanded my interests to genetically encoded sensors of cellular dynamics. As I began learning about various types of imaging sensors, calcium ion (Ca2+) indicators, in particular, stuck out to me due to their variety and depth of development. As I am unable to take part in in-person projects due to COVID restrictions, to ensure my understanding of the topics I was reading about, I began to write this review.
Abstract
Methods of performing neuroscience research have progressed remarkably in recent years, providing answers to many different types of questions. Genetically encoded indicators (sensors) are of particular interest for use in answering questions about neural circuits, cell specific populations, and single cell dynamics. These indicators modulate their fluorescence in different cellular environments and allow for optical observations. Of the various cellular activities that can be measured by genetically encoded indicators, the dynamics of calcium ions (Ca2+) are of interest due to their fundamental importance in neuronal signaling. In this review, we introduce the basic underlying features of genetically encoded calcium indicators (GECIs) including characteristics of fluorescence, an overview of GECI engineering, and a brief discussion of some common variants of GECIs and their uses.
Introduction
Modern neuroscientists have found many ways to analyze the information-carrying neuronal circuits and dynamics within the brain. The continued development of genetically encoded optical indicators, specifically Ca2+ indicators, is particularly promising for analyses of single neurons or neural circuits. Optical fluorescence imaging allows large populations of neurons to be examined simultaneously and avoids major damage to the cells of interest [1]. In particular, measuring the dynamics of Ca2+ can be useful in inferring spiking activity in neurons, as Ca2+ is involved in neuronal action potentials. In this review, we will introduce the basic workings of genetically encoded sensors beginning with a ground-up introduction of fluorescence measurements and the process of engineering genetically encoded sensors. Several Ca2+ indicators will be briefly discussed in order to examine recent progress and how this can impact future studies of the brain.
Mechanics of Fluorescence Indicators
Fluorescence imaging is a valuable tool for visualizing populations of cells. It is relatively non-invasive; but, in order to use optical tools to study the cortex, some surgical procedures must still be performed. A cranial window might be installed in the animal to shine light through; alternatively, an endoscope or fiber optic cable could be installed at the desired depth within brain tissue [2].
Fluorescent proteins (FPs) internally form a barrel-like structure containing the chromophore (also known as the fluorophore), which is a trio of amino acids responsible for the protein’s fluorescence. The chromophore is autocatalytically formed as a post-translational modification, requiring just atmospheric oxygen. Genetically encoded indicators rely on a change in the chromophore environment within the labeling FP. Fluorescence is observed when light of an appropriate wavelength excites the chromophore’s electrons, which then results in the emission of a lower energy photon as the excited electrons return to a lower energy state. FPs are often connected to sensing domains, which induce the change in the chromophore environment after detecting the event of interest (e.g., Ca2+ binding for GECIs). Any number of cellular activities may induce a conformational change such as changes in pH or the binding of a ligand. Sensors can have one or two FPs with partially overlapping fluorescence spectra. Single FP-based sensors are generally preferred as indicators; the green fluorescent protein (GFP), cloned from the jellyfish Aequorea Victoria, is the most commonly used FP for single FP sensors [3]. In systems with two FPs, a Förster or fluorescence resonance energy transfer (FRET) occurs. FRET involves an energy transfer from the higher energy (more blue-shifted) donor FP to the lower energy (more red-shifted) acceptor FP. Genetically manipulating FP systems by circularly permuting FPs (fusing the original termini of the FP and introducing a new opening closer to the chromophore) can improve their performance in sensors by making the chromophore more accessible to the outside environment and, thus, more susceptible to environmental changes [4]. FPs like GFPs are also typically oligomeric in their natural environment (i.e. multiple copies stick together); but, in order to help prevent breakdown and allow for better combination with sensing domains in indicators, FPs must also be mutated to become monomeric [5].
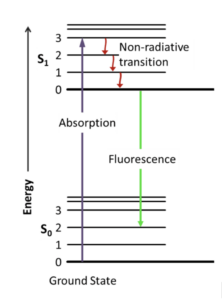
Figure 1: A Jablonski diagram that visualizes an electron’s excitation to a higher energy level by absorption of a photon and subsequent fluorescence emission with energy decay.
To image fluorescent systems, we can use fluorescence microscopy with one or multiple photons (Fig. 2) [2]. In one-photon systems, the fluorophore absorbs energy from a light source and is excited by a single photon. Some energy is lost non-radiatively (without light) resulting in the emittance of lower energy, visible photons from the fluorophore. One-photon systems are relatively inexpensive and fast but can only penetrate tissue to a shallow depth. In contrast, multi-photon microscopy shows more promise for in vivo imaging because of its reduced out of focus emission, light scattering, and phototoxicity. The combination of the energy of multiple photons is required for excitation in such systems.
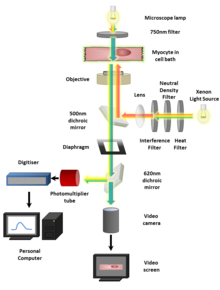
Figure 2: A diagram outlining the setup for a standard fluorescence microscopy experiment.
Genetic encoding of sensors
To introduce indicator genes into a system, methods like in utero electroporation or viral vectors can be used [6, 7]. DNA promoters or localization sequences can be used to target specific subtypes of neurons in organisms to produce transgenic animals. Transgenic animal genomes that have been modified by artificial bacterial chromosomes, CRISPR, or effector nucleases, and are particularly useful when longitudinal and intensive sampling is required. Genetic changes can be maintained throughout an animal’s lifespan and lines of transgenic animals can be bred for further testing [2]. A recombinase system administered via viral vector, like the popular Cre/loxP system, can be used to achieve high specificity [6]. In the Cre/loxP system, the loxP sequences are placed at specific target sites of genomic DNA. The Cre-recombinase protein can then target loxP sequences to modify the genetic sequence. Two mouse lines, one carrying the gene of interest flanked by loxP sequences, and the other line expressing Cre-recombinase, can be bred to produce mice expressing the gene of interest. The Cre Driver mouse line expressing Cre-recombinase can be designed to only express the gene under certain conditions. To apply Cre/loxP to genetically encoded indicator systems, a viral vector injects the indicator genes into the brain cells of a Cre Driver mouse. The indicator is only expressed where the Cre-recombinase is active. Expression would continue through one animal’s lifetime; to create a line of mice that express the desired indicator, other methods must be used [6]. Through recombinase methods, the development of transgenic animal lines is an area of active improvement.
There are several advantages to genetically encoding indicators over other methods of imaging. There are a wide variety of neuronal events that can be observed by constructing indicators from proteins that respond to cellular events, including changes in neurotransmitter concentrations, transmembrane voltage, Ca2+ dynamics, and pH [1]. Genetic encoding also allows for selective sampling of cells based on genotype. Selective sampling is not possible with chemical dyes, nor is the viewing of the evolution of neuronal dynamics during learning or development processes [8, 1]. Similar to chemical dyes, genetically encoded indicators allow for the imaging of brain activity in neurons in vitro and in animals [9]. Neurons have the machinery implanted within them to automatically report cellular dynamics of interest.
There are several different broad classes of genetically encoded indicators that are based on the dynamics of the action potential [7]. Genetically encoded voltage indicators (GEVIs) operate based on the membrane depolarization that occurs during action potentials. Other indicators, like pH and neurotransmitter sensors, detect vesicular release. Genetically encoded pH sensors (GEPIs) react to the decrease in acidity as vesicles fuse with the membrane, and genetically encoded transmitter indicators (GETIs) are used to visualize the release of neurotransmitters into the synapse [1]. Genetically encoded calcium indicators (GECIs) operate based on the rise in cytosolic Ca2+ during an action potential; however, they do not directly measure spiking activity. When an action potential occurs, Ca2+ floods into the cell. Ca2+ influx is important because calcium ions are crucial for the release of neurotransmitters from vesicles, which then go on to produce signals in other neurons. More mild calcium ion dynamics are always present in neurons, even in a resting state. Among these various classes, GECIs have been perhaps some of the most developed of these indicators and, thus, some of the most promising.
Engineering genetically-encoded calcium indicators
Performance Criteria
As GECIs are engineered, many performance criteria must be considered. Tradeoffs often occur between the various important qualities of an indicator’s performance [10]. As we optimize the indicator to produce a desired result in one criterion, another criterion often decreases in quality. Thus, development of sensors optimized for specific applications is continuous. Some of these criteria are affinity, sensitivity, kinetics, localization, and photophysical characteristics.
Affinity, represented as the dissociation constant Kd, describes what percentage of the indicator is unbound given a particular concentration of ligand.
Specificity refers to the indicator system’s ability to respond only to the target of interest, as opposed to perhaps similar molecules.
Sensitivity is usually represented by ΔF/F0, the fractional fluorescence change, which is the fluorescence signal change over a change in concentration of the target molecule. It can also be represented by signal-to-noise ratio (SNR), the relative difference between the signal of interest and background noise.
Kinetics is the rate of change in fluorescence intensity of the indicator in response to the change in ligand concentration. There tends to be a tradeoff between affinity and kinetics [8].
Photophysical qualities like brightness, photostability, and photoswitching behaviors are also important considerations. In general, brighter or more intensely emitting indicators are desired. Photostability is inversely proportional to the rate of photobleaching (the damaging of the FP so that it becomes unable to fluoresce). Additionally, some indicators have broader ranges of excitation than others, or may change their intensity or sensitivity in different light conditions, which would limit usage.
GECIs are some of the most widely used genetically encoded indicators in vivo because of their relatively high SNR and improved properties like brightness, photostability, and dynamic range [2]. However, there are still numerous obstacles to be faced in designing GECIS, and only certain variants have faced success in vivo.
Engineering GECIs
Genetically encoded indicators generally are composed of an analyte-binding (sensing domain) and a fluorescent protein (reporting domain), though there are additional peptide complexes that assist in changing the conformation of the system [2]. Upon the occurrence of a sensing event, the sensing domain undergoes a conformational change which, in turn, induces a conformational change in the FP, resulting in fluorescent activity. Engineers of GECIs use two different strategies for constructing reporting domains: FRET-based indicators and single FP-based indicators [7]. When Ca2+ binds to FRET-based indicators, the spatial relationship between the donor and acceptor FPs changes so that there is a transfer of energy from the donor FP to the acceptor FP [2]. One family of indicators, Cameleon, has had some success. In this family, the sensing and peptide complex is located between two FPs with overlapping spectra. FRET-based indicators’ SNR tends to be lower, meaning it is harder to isolate the activity of a neuron from background noise. Because of these drawbacks, we mostly examine the engineering of the more commonly used single FP-based GECIs.
There are two popular designs among developers of single FP GECIs [8]. One is based on one of the earliest lines of calcium indicators, GCaMP. GCaMP consists of a circularly permuted green fluorescent protein (cpGFP) inserted between the Ca2+-binding protein, calmodulin (CaM), and another peptide called RS20, which binds CaM upon Ca2+ binding. When CaM binds Ca2+, a conformational change is induced in the cpGFP and the sensor fluoresces [10, 11]. Another recent design, the NTnC family of indicators, inserts a calcium-binding domain into a split FP [8]. Unlike GCaMP-type indicators, NTnC indicators display an inverted fluorescence response upon calcium binding (i.e., fluorescence decreases upon Ca2+ binding). They are less optimized than GCaMP variants, but it is hypothesized that their lesser Ca2+ binding capacity would interfere less with normal calcium dynamics.
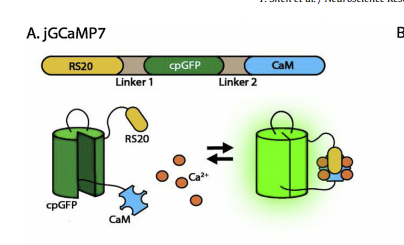
Figure 3: A basic representation of the GCaMP structure.
There have been efforts to expand the color variants of GECIs. In particular, there has been much effort to develop red-fluorescing GECIs because longer red wavelengths reduce phototoxicity and have better tissue penetration [12]. However, there have been many obstacles to producing red-fluorescing GECIs. Unlike GFP, inserting calcium binding domains into red fluorescent proteins (RFPs) disrupts folding and chromophore maturation [8]. A more popular design choice is to replace the GFP in a GCaMP-style indicator with an RFP and optimize the sensor for a new FP [1].
GECIs are improved iteratively through directed evolution and linker optimization between the cpFP and the sensing domains. Site-directed mutagenesis can be used to mutate specific locations to produce novel variants. In the development of one variant of GCaMP, mutations were specifically introduced in the calcium-binding domain-cpGFP linker in a GCaMP5 scaffold to increase sensitivity [11]. Using directed evolution, mutations are randomly introduced. Then, upon testing for desired effects, the variants that produced the best results may be preserved and propagated. This process may repeat many times, producing increasingly successful indicators as the best-performing mutations survive across generations.
Challenges
Genetically encoding indicators, as a rule, comes with challenges. If we choose to use viral infection as our genetic encoding scheme, consideration must be taken to the many viral serotypes, which have varying levels of efficiency and can be toxic. Furthermore, in utero electroporation can be unpredictable, and transgenic animals may not express indicators at sufficiently high levels to be useful [1].
Sensors may affect the natural dynamics of their measured systems, affecting accuracy of results. GECIs, particularly GCaMP-based designs, may interfere with regular Ca2+ dynamics and gene expression [8]. This interference is likely due to interactions between the calcium-binding sensing domain with native proteins and the lack of availability of calcium once bound to the indicators. There have been efforts to improve and modify the calcium-binding domain so that it can bind fewer Ca2+ or otherwise improve affinity so that the indicator operates at lower concentrations of calcium.
There is also difficulty in using these indicators in vivo [2]. Especially in the mammalian brain, the SNR is highly decreased due to the amount of background noise. This reduced SNR is putting aside the level of breakdown that naturally happens in vivo vs. conditioned, cultured environments. Although many indicators have improved structural integrity in vivo, there are many that still cannot be used in living organisms.
Progress in GFP-based GECIs
There has been much development in the GCaMP series as variants are continuously improved by site-directed mutagenesis and computational design efforts [2]. The jGCaMP7 series, built from the GCaMP-6 series, provides a good example of optimization of indicators for different purposes: jGCaMP7f is optimized for fast kinetics, jGCaMP7s is optimized for high sensitivity (though it has slower kinetics), and jGCaMP7b is optimized to have a brighter baseline fluorescence [11]. All of these indicators are based on the same base scaffold but differ drastically in performance because of just a few mutations in the CaM-binding peptide, the GFP, the CaM domain, or the linkers between domains.
Progress in RFP-based GECIs
RFP-based GECIs have important advantages over GFP-based ones. Beyond the importance of color variety in tracking distinct populations of cells at once, red GECIs are also promising for reducing phototoxicity and allowing deeper imaging [12]. There are many promising RFP-based GECIs being developed, though they are generally dimmer than GFP-based indicators and may display photobleaching behaviors under blue light [1]. In particular, there are R-GECO1 variants like jRGECO1a, the RCaMP series, and, perhaps most promising, K-GECO1 [12]. There are three widely used RFPs from which red GECIs are developed; each red indicator family was generated from different RFPs. K-GECO1 has shown particular promise as it works at a distinct spectral range, allowing researchers to simultaneously work with other indicators in multicolor imaging experiments, and it also shows minimal fluorescent noise [9].
Designs of red GECIs often expand on the GCaMP design–for example, K-GECO1 follows a similar design of sandwiching the circularly permuted FP between the Ca2+-sensing domain, CaM, and a CaM-binding peptide [12]. Switching the GFP in GCaMP with an RFP comes with engineering challenges of linker optimization and preventing the breakdown of the sensor. The increased penetrative depth of red GECIs has been used to image subcortical areas like the hippocampus or medial prefrontal cortex relatively noninvasively, demonstrating the applicability of GECIs in neuroscience research [13].
There are other FP-based GECIs in development, but of particular interest is the development of near-infrared GECIs, whose spectral distinction from other indicators would help prevent photoswitching when used with optogenetic tools [8, 14].
Uses and Applications
The applications of GECIs are varied and powerful. The use of genetically encoded indicators allows for the analysis of cells of a specific type or subpopulation as they select for specific genetic qualities. The first transgenic mouse line expressing GCaMP2 in the cerebellar cortex was generated in 2007 and has allowed for characterization of certain synapses [6]. GECIs have been used to provide single-cell resolution to the decades-long study of various topographic maps in the brain and to track the communication of neural circuits [2]. In rats, GECIs have been used to monitor neural population behavior during motor learning tasks and observe the response of cells to sensory deprivation in the primary visual cortex after retinal lesion. They allow examination of ensemble and single cell-scale neural events at more and more temporally precise levels. Broadly, and perhaps more importantly, they are often used in conjunction with optogenetic and other experimental methods that allow for the inference of causation. In using these indicators, the stimulation techniques used in optogenetic experiments can also involve precise tracking of calcium or other dynamics in cells of interest [8]. These experimental approaches have caused excitement as they allow for the examination of behaviors of cells or whole organisms upon physical stimulation of even just single cells. The continued expansion of these approaches is promising.
Conclusion
Many researchers are devoted to developing new and distinct calcium indicators based on existing indicator series. With more GECIs than ever available to neuroscientists, there is some challenge in choosing which is best suited to the exploration of a particular question. With the continuing development of mouse lines and methods of genetically encoding more potent indicators with high temporal resolution, GECIs will continue to be an increasingly important tool within the neuroscientist’s toolkit that allows for population or single-cell imaging with greater resolution than ever before.
References:
- Lin M, Schnitzer M. 2016. Genetically encoded indicators of neuronal activity. Nature Neuroscience 19(9):1142-1153.
- Broussard G, Ruqiang L, Tian L. 2014. Monitoring activity in neural circuits with genetically encoded indicators. Frontiers Molecular Neuroscience 7.
- Cranfill P, et al. 2016. Quantitative assessment of fluorescent proteins. Nature Methods 13(7):557-563.
- Baird G, Zacharias D, Tsien R. 1999. Circular permutation and receptor insertion within green fluorescent proteins. PNAS 96(20):11241-11246.
- Zacharias D, et al. 2002. Partitioning of Lipid-Modified Monomeric GFPs into Membrane Microdomains of Live Cells. Science 296:913-916.
- Knöpfel T. 2012. Genetically encoded optical indicators for the analysis of neuronal circuits. Nature Neuroscience. 13:687-700.
- Wang W, Kim C, Ting A. 2019. Molecular tools for imaging and recording neuronal activity. Nature Chemical Biology 15:101-110.
- Piatkevich K, Murdock M, Subach Fedor. 2019. Advances in Engineering and Application of Optogenetic Indicators for Neuroscience. Applied Sciences 9(3):562.
- Shen Y, et al. 2018. A genetically encoded Ca2+ indicator based on circularly permutated sea anemone red fluorescent protein eqFP578. BMC Biology 16(9).
- Shen Y, et al. 2020. Engineering genetically encoded fluorescent indicators for imaging of neuronal activity: Progress and prospects. Neuroscience Research 152:3-14.
- Dana H, et al. 2019. High-performance calcium sensors for imaging activity in neuronal populations and microcompartments. Nature Methods 16:649-657.
- Molina R, et al. 2019. Understanding the Fluorescence Change in Red Genetically Encoded Calcium Ion Indicators. Biophysical Journal 116:1873-1886.
- Kondo, Masashi, et al. 2017. Two-photon calcium imaging of the medial prefrontal cortex and hippocampus without cortical invasion. eLife Neuroscience.
- Qian Y, et al. 2019. A genetically encoded near-infrared fluorescent calcium ion indicator. Nature Methods 16:171-174.

