By Saloni Dhopte, Genetics and Genomics, ’23
Author’s note: Do you think CRISPR-Cas9 genome editing is amazing? Well, let me tell you about another technique that has been proven to be more accurate and efficient than CRISPR systems. It’s prime editing – a method of genome editing that utilizes a single stranded nick to edit DNA! I first learnt about prime editing from my then graduate student mentor, Dr. Peter Lynagh, while I was working in the Comai Lab at UC Davis. I remember scouring the internet for papers and looking up applications of this revolutionary technology, as it fascinated me beyond measure. So naturally, when we were tasked with writing a scientific review for my UWP 102B class, I had to write about this! Prime editing is an up-and-coming tool in gene editing, and I hope through my review, readers with basic to intermediate understanding of molecular genetics are able to share my amazement and admiration for prime editing!
Abstract
Current methods of genome editing involve a double stranded break on the DNA molecule, thus involving high frequency of unwanted base pair insertion/deletions (indels) and off-target effects in the repair process. Some are relatively limited in scope in terms of the length of the edit and specific type of base pair substitution. Recently, a new technique called prime editing has been developed which creates a single stranded break and shows high accuracy and editing proficiency along with minimal off-target effects in the genome. The molecular machinery of prime editing is very accurate; it recognizes the region of interest in the genome and is precisely able to insert an edited DNA sequence. In this review we discuss the mechanics of prime editing, compare it to other methods of genome editing, and investigate its applications as well as limitations in the health field. Numerous papers from the PubMed database point towards the potential of prime editing in repairing disease-causing mutations in humans. However, researchers are not using it for in vivo experiments (experiments that take place in a whole living organism) just yet, as they believe we need to learn a lot more about safe methods of delivery to human cell lines, side effects of the treatment, and overall efficiency of editing. Regardless, a lot of progress is consistently being made, including optimization of the prime editing machinery, online search databases, and ex-vivo applications. With the current pace of scientific discovery, the goal of using prime editing to its full potential will become reality soon enough.
Keywords: single-stranded break, genome editing, CRISPR-Cas9, guideRNA, disease-causing mutation repair.
Introduction
In 2019, Dr. David Liu published a paper introducing a new method of genome editing called prime editing (PE). The method is novel in its approach since it surpasses a majority of the drawbacks of pre-existing methods of genome engineering such as base editing (BE) and Clusters of Regularly Interspaced Palindromic Repeats (CRISPR)-CRISPR associated protein (Cas9), a method of genome editing adapted from the bacterial immune system’s defense mechanism. The main reason why PE is able to overcome these drawbacks is because it involves a single stranded break on the DNA molecule, as opposed to CRISPR-Cas9 which involves a double-stranded break (DSB), thus significantly reducing unwanted indels in the genome [1]. So far, a lot of studies have compared the efficiencies of PE with BE and CRISPR-Cas9. PE has been put to use in modeling diseases in organoids (tissue cultures that mimic in vivo organs) and researchers are developing ways of correcting the mutations that cause these diseases. In theory, PE could correct 90% of all disease-causing mutations in humans [2]. However, the unanimous opinion remains: a lot of work needs to be done in the field before PE can be used to correct mutations in a safe manner in vivo [1], [3], [4].
In this review I will investigate the mechanism and scope of prime editing and see how it can be used to study disease-causing mutations. By analyzing the technique and comparing it against the existing and well researched types of genome-editing, I will investigate PE’s effectiveness in repairing these mutations and see how the mechanism can be optimized.
How does Prime Editing work?
Mechanism
The prime editing system is based off of the CRISPR-Cas9 genome editing system. In CRISPR-Cas9, there is a single-stranded, guide RNA molecule (sgRNA) that is complementary to a specific region of the DNA and it is associated with a DNA endonuclease enzyme, Cas9. The sgRNA searches for and binds to the sequence homologies within the genome. Directed by the sgRNA to this complementary region of interest, the Cas9 protein then cuts the DNA, creating a DSB[2]. This DSB can then be repaired by double-strand break repair processes of varying levels of efficiency. In the case of prime editing, we still have the guide RNA, but it is modified such that it includes the sequence of our desired edit (referred to as prime editing guide RNA or pegRNA). Further, the Cas9 protein is partially inactivated (referred to as Cas9-nickase) such that it only cuts the 3’ strand of the complementary region on the DNA- creating a single stranded break [5]. And this makes all the difference. Since PE involves a single stranded break, the machinery doesn’t rely on the inefficient and unpredictable DSB repair mechanisms.
The single stranded break caused by Cas9 nickase leads to the formation of a 3’ DNA flap. This flap binds to a sequence on the pegRNA called the primer binding site (PBS). The reverse transcriptase enzyme, which is fused to Cas9 nickase, elongates the DNA using the 3’ flap as a primer, thus synthesizing a new “edited” DNA strand. The newly edited strand can then be incorporated into the original DNA molecule through a number of ways. The various approaches through which this resolution happens makes up the different kinds of prime editing.
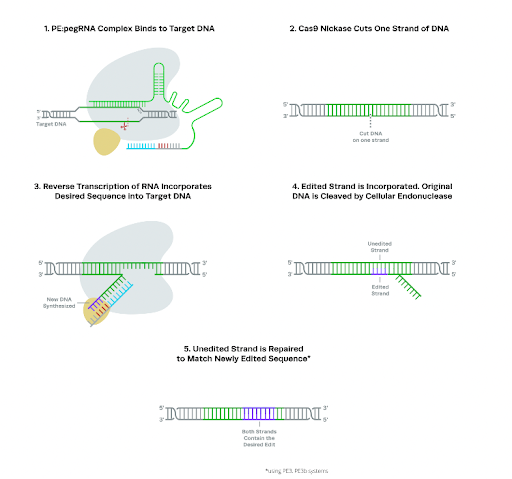
Reproducible Graphic : The mechanics of prime editing
https://www.synthego.com/guide/crispr-methods/prime-editing
Types of Prime Editing
The mechanism PE1 relies on an endogenous endonuclease – FEN1, which cuts the 5’ end of our unedited DNA so that it doesn’t come in the way of our edited strand. With this cut, our edited DNA is thermodynamically favored to bind to the original DNA molecule. For PE2, the Liu group modified the sequence of RT to increase its thermostability, binding to DNA template and enzyme processivity. To take the mechanism to the next level, Anzalone et al. developed PE3 where they introduced a second guide RNA (gRNA) molecule that creates a 5’ nick on the non-edited strand, which allows for the edited DNA strand to be used as a template to complete the process. Finally, we have PE3b in which the gRNA is programmed such that it creates the 5’ nick only after the edited strand has been completely formed [5]. This approach led to reduced indel formation and improved editing. According to Anzalone et al., PE3 is 1.5 to 4.2-fold more efficient than PE2. PE3b, while not showing a significantly higher editing efficiency, has a 13-fold reduction in indel formation [2].
To summarize, the way PE works is by utilizing a special RNA molecule called pegRNA that encodes our desired DNA edit and targets the exact region of DNA where the edit has to be incorporated. Along with the Cas-9 nickase, it accurately creates a single stranded break, synthesizes the edited DNA, and finally incorporates the edit through various mechanisms – PE1, PE2, PE3 and PE3b.
Why is Prime Editing the best genome editing method we know?
Prime Editing versus CRISPR-Cas9
There are multiple characteristics of PE that make it better than our pre-existing methods of genome editing.
First, PE involves a single stranded break in the DNA as opposed to CRISPR-Cas9 editing which involves a double stranded break (DSB). A DSB can be repaired in one of two ways – via homology directed repair (HDR) or non-homologous end joining (NHEJ). The latter leads to a lot of indels. Thus, the editing efficiency of genome editing methods employing DSB’s is low. In contrast, PE doesn’t involve a DSB to begin with, thus the process of repairing the break doesn’t rely on the generation of random indels. Chemello et. al showed how the mutations causing Duchenne Muscular Dystrophy (DMD) are corrected with less unwanted effects via PE as opposed to CRISPR-Cas9 [4].They used prime editing to correct one of the most common mutations of DMD – the deletion of exon 51. Chemello et al. first attempted to restore the correct open reading frame (ORF) by inducing exon skipping. They used CRISPR Cas9 to systematically make two cuts such that exon 52 would be skipped, and the ORF would be restored. However they were unsuccessful due to the high rate of indels and off-target effects of CRISPR-Cas9. On the other hand, with PE, they were able to reframe the exon and precisely inserted two nucleotides into exon 52, thus bypassing the need for exon skipping entirely. This demonstrates the ability of PE to specifically target and edit DNA sequences in order to correct disease-causing mutations without the unwanted effects of double-stranded break repair pathways.
Further, since PE requires 3 separate hybridization events (pegRNA spacer to target DNA for Cas9 binding, pegRNA PBS to target DNA, and target DNA 3’ flap to RT product) to occur, it has significantly less off-target effects in the genome. In CRISPR-Cas9, the guide RNA, as it is searching DNA for complementarity, can bind to other regions of the genome with similar sequences, leading to DSB’s in places that were not targeted [2]. Kim et al. were unsuccessful in trying to correct DMD using CRISPR-Cas9, due to these off-target effects. However, they found no significant unwanted indels in the genome of mice hepatocytes that were prime edited to correct for HT1 [6]. Similarly, Geurts et al. performed whole genome sequence analysis on prime edited five colon organoids and reported no mutational differences among the edited organoid sequences [3]. These findings establish the safety of PE as compared to CRISPR-Cas9.
Prime Editing versus base editing
PE’s battle with base editing is not as one-sided as it is with CRISPR Cas9. Base editing performs rather well in most experiments.
There are two kinds of base editing – adenine base editing that can change a nucleotide from A to G or G to A, and cytosine base editing, which changes C to T or T to C. The mechanism involves an inactive Cas9 protein (dCas9) fused to a deaminase molecule which makes the respective base change possible [1]. Base editing doesn’t involve a break on the DNA molecule at all. It also doesn’t rely on the generation of random indels for editing. Hence, it is not surprising that the efficiency of this method surpasses that of PE. Geurts et al. reported that base editing induced correct mutations in 50% of the colonic organoids whereas prime editing was only able to reach 22% [3]. Similarly, Schene et al. found that using PE for the correction of mutations in liver organoids was less effective than using base editing. In this way, both papers report the same finding – when working with a mutation that can be corrected by base editing, it outperforms PE.
But here is the catch. First, base editing can make only four of the twelve possible base pair changes. If the disease of interest requires an adenine to be corrected into a cytosine, base editing doesn’t even come into the picture– that substitution is beyond its capability. This severely limits the scope of genome editing. The second drawback deals with the size of the edit. Some diseases require a stretch of nucleotides to be corrected– not just a single nucleotide. If the editing window for BE is increased to more than one nucleotide, especially if the edit includes more A or C bases, a lot of by-stander edits are observed. This is because the Cas9-deaminase complex makes all base substitutions in its range, including those we don’t want [2]. This problem doesn’t arise in PE because the pegRNA encodes highly specific DNA insertions up to 80 base pairs in length. Because 98-99% of insertions, deletions and duplications in the pathogenic human genetic variants are smaller than 30 base pairs, researchers have claimed that with PE, we will be able to correct 90% of disease-causing mutations in humans [1].
What has been accomplished so far using Prime Editing?
One of the most sought-after goals of genome editing is to be able to correct diseases-causing mutations. While PE is being used increasingly for its precision in editing DNA, it is a relatively new technique and so all of the research takes place in vitro ( in an artificial environment simulated to mimic the human body). In the hopes of eventually overcoming this gap and moving on to in vivo studies, scientists are also working on optimizing PE to have even less off-target effects. They have varied the molecular machinery, model organoids that mimic in vivo organ systems, and target mutations in different combinations, to find an approach with the best results. Optimization of the prime editing machinery is a well-established path to achieving the goal of using this technique in therapeutic applications.
Repair and modeling of disease-causing variants
Prime editing has been successfully used to model several diseases in human organoids. Broadly speaking, there are two main goals of these particular studies. First, to study the efficiency of PE, and working on ways of improving the mechanism. Geurts et al. utilized PE to model the mutation causing cystic fibrosis in human adult derived colonic organoids and then used PE to correct the mutation. Scientists employed both PE and BE for these steps and found that BE was more efficient in inducing intended mutations as compared to PE, but again, it remains limited to 4 of the 12 base pair substitutions. They acknowledged that if edits need to be made outside of this window, PE is the best approach [3].
The second kind of PE editing studies investigate the ways in which disease-causing mutations can be corrected. Schene et al. modeled mutations in liver organoids to mimic the development of liver cancer, then used PE [1]. Schene et al. and Guerts et al. both confirmed that PE is the better choice only for the subset of mutations not applicable for correction using BE [3].
With that, it is quite clear that we need to work a lot more on PE to further increase its efficiency. To do this, multiple researchers are focusing their efforts on optimization of PE.
Optimization of pegRNA’s
The prime editing machinery is highly advanced in structure. Compared to CRISPR-Cas9, there are fewer elements involved and thus less unwanted indels. The key player making this possible is the prime editing guide RNA (pegRNA). Researchers have worked extensively on optimizing the performance of the pegRNA through a variety of approaches. Lin et al. identified two main factors that have shown increased efficiency of editing: first, designing primer binding sites (PBS) on the pegRNA with melting temperature less than 30 degrees Celsius, and second: using not one, but twopegRNA’s encoding the same edit [7]. Together, these boost the editing efficiency 17.4-fold.
Moreover, there is an increase in resources and tools for pegRNA optimization. PegFinder is an online software that allows scientists to program the specific pegRNA to fit their experiment [3]. More recently, Lin et al describe the construction of their own web application called PlantPegDesigner. They claim that their tool is more user-friendly than PegFinder, as the latter necessitates experimental testing of pegRNA’s. PlantPegDesigner only requires a single DNA sequence as an input and provides a variety of parameters to be optimized by the user – an ideal candidate pegRNA [7]. This technology has the potential to greatly simplify prime editing experiments with plants, which in turn might lead to quickly reducing the knowledge gap in the field. Another similar web application is PrimeDesign – a tool that not only provides the user with an ideal pegRNA but visualizes the entire prime editing event. It allows users to rank pegRNA’s based on efficiency and includes extensive annotations. Additionally, Hsu et al. created a database called PrimeVar using all of these results, which can be used to search for pegRNA’s correcting ~70,000 pathogenic human genetic variants [8].
Conclusion
Prime editing is a novel breakthrough in the field of genome editing. It has been only three years since the publishing of Dr. David Liu’s original paper introducing the world to PE. PE is able to target and edit any region of the genome while avoiding drawbacks of current gene-editing methods, made possible by the induction of a single stranded break. Scientists have demonstrated the superiority of PE when compared to base editing and CRISPR-Cas9 editing. BE, although more accurate and known for less off-target effects than PE, can only correct a subset of base-pair substitutions. CRISPR-Cas9 involves a double stranded break on the DNA molecule, leading to high rates of unwanted insertions/deletions in the genome as compared to PE. Within a span of two years, four distinct types of PE have been developed – PE1, PE2, PE3 and PE4 – each more efficient than the last. The development of online tools such as PrimeDesign and PlantPegDesigner show the rate at which scientists are making progress with PE. However, we are far from the finish line. Most researchers still remain skeptical about the use of PE for in-vivo applications. While some say it is imperative to develop safe methods of delivery to human cell lines, others question the consequences of off-target effects in the genome. We don’t fully understand how PE might affect other cells of the subject [2]. Additionally, researchers aren’t certain about the longevity of prime edited disease corrections [4]. Most agree that in theory, prime editing will be revolutionary in terms of advancing human health, but given the relative recentness of the technology, there is still a lot of work to be done. Despite the gray area, PE certainly has a lot of potential and will be one of our strongest tools in improving human health in the future.
References:
- I. F. Schene et al., “Prime editing for functional repair in patient-derived disease models,” Nat Commun, vol. 11, no. 1, p. 5352, Dec. 2020, doi: 10.1038/s41467-020-19136-7.
- A. V. Anzalone et al., “Search-and-replace genome editing without double-strand breaks or donor DNA,” Nature, vol. 576, no. 7785, pp. 149–157, Dec. 2019, doi: 10.1038/s41586-019-1711-4.
- M. H. Geurts et al., “Evaluating CRISPR-based prime editing for cancer modeling and CFTR repair in organoids,” Life Sci. Alliance, vol. 4, no. 10, p. e202000940, Oct. 2021, doi: 10.26508/lsa.202000940.
- F. Chemello et al., “Precise correction of Duchenne muscular dystrophy exon deletion mutations by base and prime editing,” Sci. Adv., vol. 7, no. 18, p. eabg4910, Apr. 2021, doi: 10.1126/sciadv.abg4910.
- H. Jang et al., “Application of prime editing to the correction of mutations and phenotypes in adult mice with liver and eye diseases,” Nat Biomed Eng, vol. 6, no. 2, pp. 181–194, Feb. 2022, doi: 10.1038/s41551-021-00788-9.
- Y. Kim et al., “Adenine base editing and prime editing of chemically derived hepatic progenitors rescue genetic liver disease,” Cell Stem Cell, vol. 28, no. 9, pp. 1614-1624.e5, Sep. 2021, doi: 10.1016/j.stem.2021.04.010.
- Q. Lin et al., “High-efficiency prime editing with optimized, paired pegRNAs in plants,” Nat Biotechnol, vol. 39, no. 8, pp. 923–927, Aug. 2021, doi: 10.1038/s41587-021-00868-w.
- J. Y. Hsu et al., “PrimeDesign software for rapid and simplified design of prime editing guide RNAs,” Nat Commun, vol. 12, no. 1, p. 1034, Dec. 2021, doi: 10.1038/s41467-021-21337-7.

